TMEM260 revealed
May 22, 2022One more human glycosyltransferase
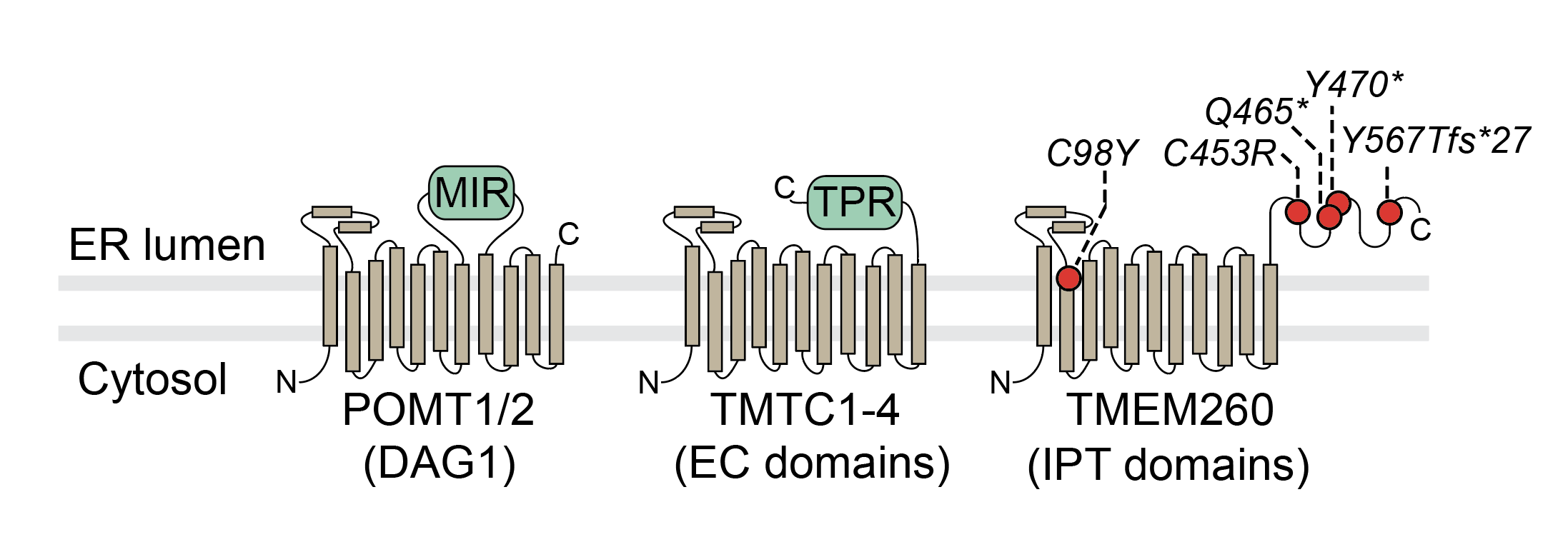
Figure 1 from the TMEM260 manuscript
The moment where I found out that we’d guessed correctly again1
We’ve found a gene, encoded for by most animals that we think is another
mannosyltransferase. For a protein glycosylation pathway that we were pretty
sure involved two enzymes, and one major substrate only a few years ago, it’s
now taking up a fair amount of space on our pathway figures. The name of the
gene is
TMEM260. Unlike the other new enzymes that we found,
it doesn’t look like there’s an opportunity to backronym2
meaning into this name, so there’s likely a renaming due for this at some point.
You can read about it at our preprint
3
that we dropped at Research Square paper
4
in PNAS. I’ll give some background information here
that didn’t sit properly in the manuscript.
Lots of guilt by association for heart defects
Soon after we decided to try and figure out if TMEM260 encoded for a glycosyltransferase, a publication came out reporting on mutations in this gene in patients with developmental defects 5 in the kidney, heart and brain. In our paper, we’ve got a lot of new data that sheds some more light onto these phenotypes, but there’s also many individual data points that also point us in that direction. We know from our glycoproteomics that this mannosyltransferase targets Plexin D1, and there’s a match between one of the phenotypes (“Truncus arteriosus”) and phenotypes arising from a deficiency in PLXND1 6 . If you dig into the phenotypes for CDG-Ie, and PMM2-CDG, you’ll find descriptions of heart defects (patent ductus arteriosus and truncus arteriosus) 7 8 9 (via 10 ). Looking upstream from the plexins,
GATA6 has been implied 11 to regulate semaphorin/plexin signalling via regulation of Plexin A2 and SEMA3C.
Beyond these more well supported phenotypes, there’s very sparse information
from when TMEM260 was known by it’s street name of C14orf101, that links it to
non-Hodgkin lymphoma survival
12
.
Apart from some reports that semaphorin signalling is implicated in adhesive and
metastatic potential, there’s little to go on.
AlphaFold2 model for TMEM260. The transmembrane region looks a lot like a standard GT-C fold, but has a C-terminal region that is unlike other Dol-P-Man mannosyltransferases ( e.g., POMT1, POMT2,
A strange C-terminal domain
One confusing thing when we first started to look at this protein was that we didn’t understand what was happening at the C-terminal. There weren’t any domains annotated for it on InterPro, and it seemed pretty unlikely that there was a good 300 amino acids of protein just sitting there without any structure.
It was a stroke of luck that I ran across the preprint for Max Bileschi’s new PFAM-N method 13 that makes use of deep learning to perform better PFAM annotations. When I dug through the data, I saw that this method finally classified the human TMTCs as part of the right PFAM family. I reached out to Max over Twitter, and we began to discuss the mannosyltransferases. I hoped that the new method would be able to identify any domains on this C-terminal. In the end, there weren’t any matches for the majority of the C-terminal domain, but he did confirm that there were bits of this region that look like TPRs. The Alphafold2 model, backed up my hunch that was something there. It’s likely that this protein will get classified into a new CAZy family, so there’ll be two brand new human families to our name!
Mostly conserved in metazoa
We can find this gene pretty broadly in eukaryotes, but interestingly, there doesn’t look to be an orthologue in the invertebrates. The conserved domain in TMEM260 is also found in prokaryotes, although it’s unclear what this enzyme would be doing in these organisms.
The future
There’s a lot of function to unpack with this new enzyme, and the fact that it glycosylates many important molecules apart from plexins (including Hepatocyte growth factor receptor and Macrophage-stimulating protein receptor) means that it’s probably going to be pretty interesting.
Footnotes
-
I happened to be down in Australia when Adnan dropped into my DMs to deliver one of the better types of message to get over DM. ↩︎
-
I’m impressed with whoever decided to rename the TMTC family of genes so that they could keep their HGNC name, but could change the meaning of the acronym from TransMembrane and TetratriCopeptide to Transmembrane O-Mannosyltransferase Targeting Cadherins. ↩︎
-
Ta-Shma et al., The American Journal of Human Genetics, 2017 ↩︎